About
Hi, I am Thomas Höfler!

I am a molecular microbiologist who studies the evolutionary and molecular genetics of viral infections. Using experimental evolution, I focus on key phenotypes such as antiviral resistance, immune evasion, virulence, as well as genetic conflict and cooperation within viral populations.
Herpesviruses are among the most prevalent and diverse animal viruses known to humanity. Virtually every animal species harbors one or more species of herpesvirus, regardless of whether they are mammals, fish, amphibians, molluscs, birds, or reptiles. Given this immense variation, herpesviruses are an ideal order for studying host-pathogen adaptation.
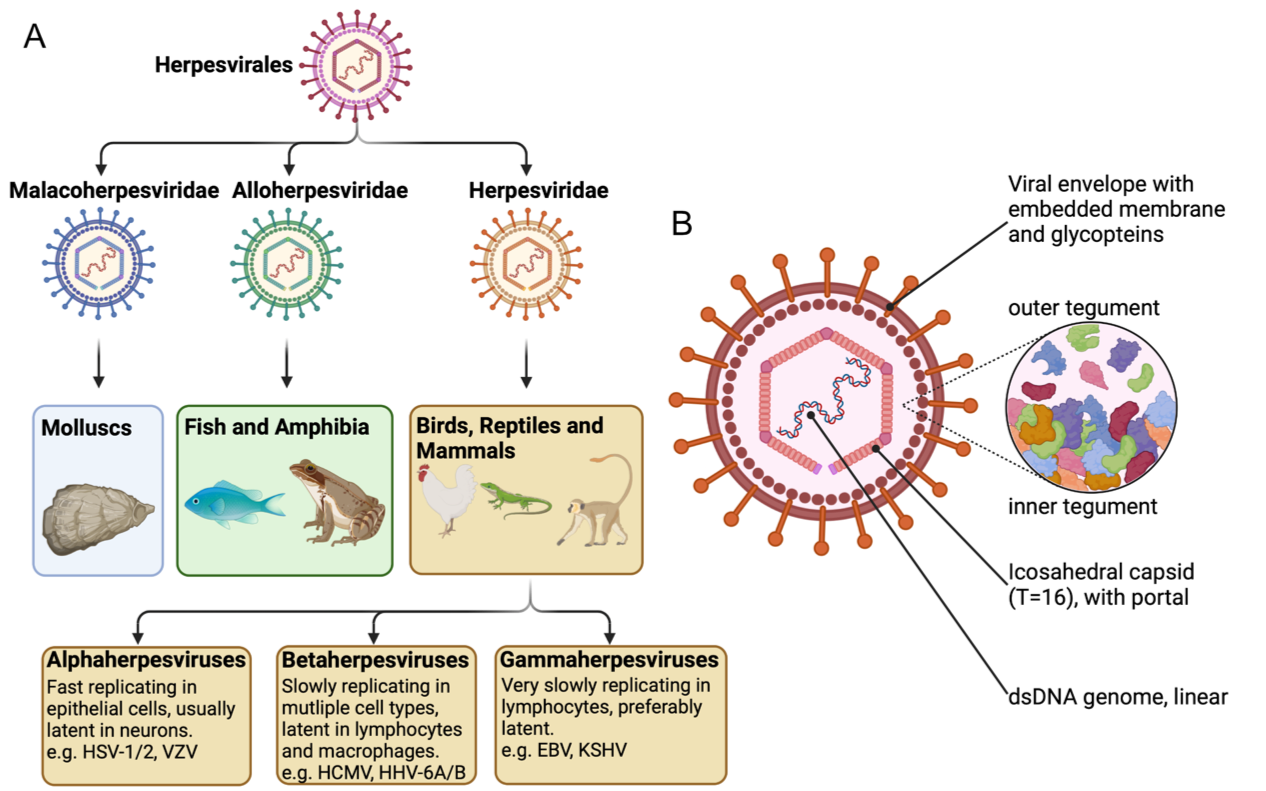
During replication of their genome, herpesviruses introduce a moderate amount of mutations. We utilized DNA-polymerase mutants to increase mutation rates and therefore accelerate evolution. We can achieve this thanks to the two separate enzymatic functions of B-family DNA-polymerases: 5’-3’ polymerase and 3’-5’ exonuclease function.
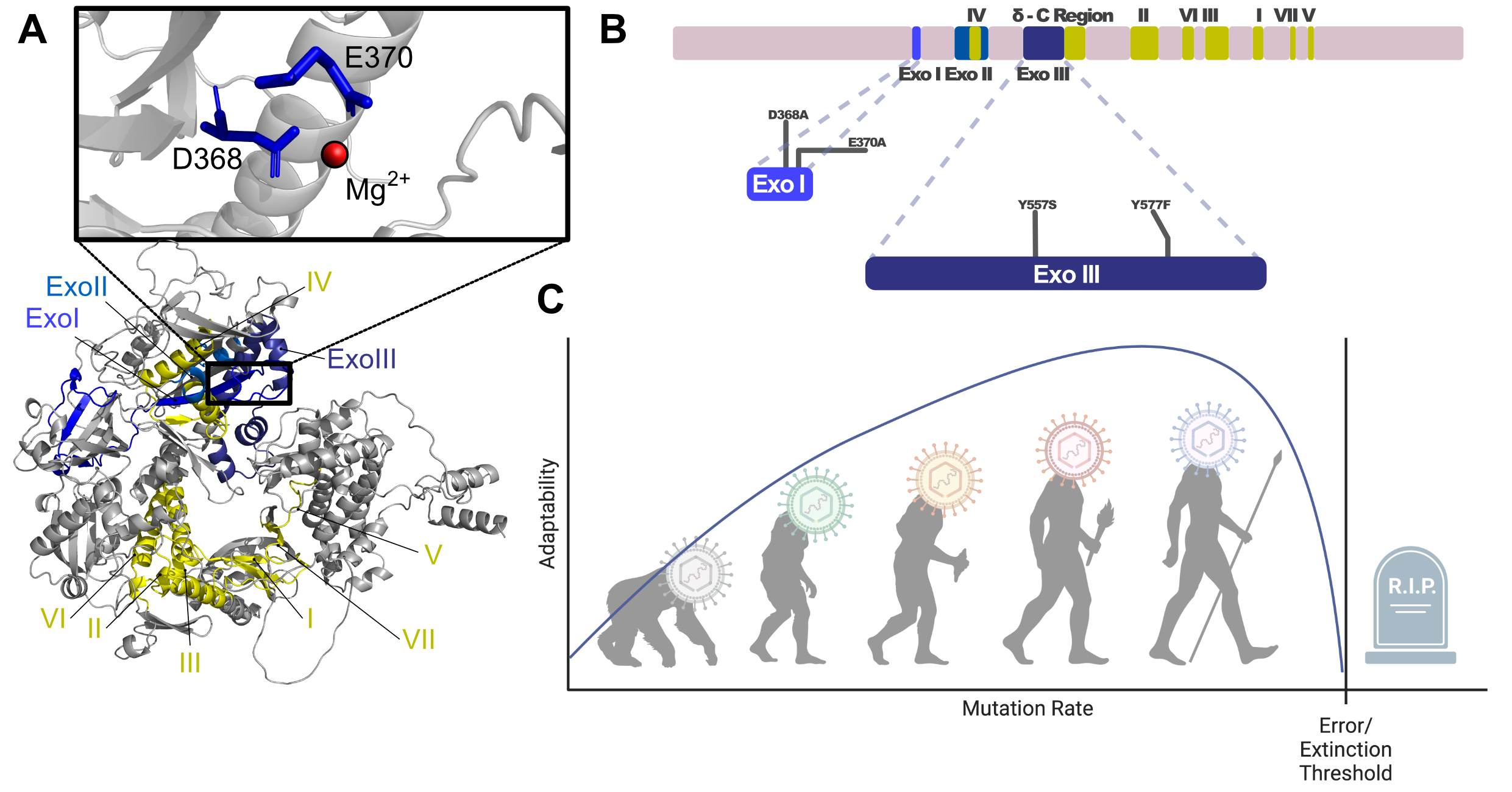
Viruses
Apart from herpesviruses, I also work on RNA viruses; here is a short list of all the viruses we handle.
- DNA viruses
- Herpes Simplex Virus Type 1 (Human)
- Human Cytomegalovirus (Human)
- Marek’s Disease Virus (Chicken)
- Channel Catfish Virus (Channel Catfish)
- Frog Virus 3 (Frogs)
- Varicella Zoster Virus (Human)
- RNA viruses
- Influenza A Virus (Mammals, Birds)
- SARS-CoV-2 (Mammals)
- West Nile Virus (Arthropods, Birds, Mammals)
- Blue tongue Virus (Midges, Cattle)
Methods
I combine wet lab bench experimentation with computational methods to build new hypotheses and confirm or reject them experimentally. Especially, genomic and transcriptomic data are integrated into AI-models to search for underlying structure and information. Both short and long-read sequencing enable an extensive exploration of genomic sequence, structure, and modification.
Publications
f you are interested, check out any of my publications; there are also additional details in the specific blog posts.
- Höfler T, Zeitlow M, Kim JY, Wyler E, Trimpert J. 2025. Rapid glycoprotein evolution enables variant interactions in herpes simplex virus type 1. Virus Evolution 11:veaf072.
- Friedrich VD, Pennitz P, Wyler E, Adler JM, Postmus D, Müller K, Teixeira Alves LG, Prigann J, Pott F, Vladimirova D, Hoefler T, Goekeri C, Landthaler M, Goffinet C, Saliba A-E, Scholz M, Witzenrath M, Trimpert J, Kirsten H, Nouailles G. 2024. Neural network-assisted humanisation of COVID-19 hamster transcriptomic data reveals matching severity states in human disease. eBioMedicine 108:105312.
- Höfler T, Nascimento MM, Zeitlow M, Kim JY, Trimpert J. 2024. Evolutionary Dynamics of Accelerated Antiviral Resistance Development in Hypermutator Herpesvirus. Molecular Biology and Evolution 41.
- Leeks A, Bono LM, Ampolini EA, Souza LS, Höfler T, Mattson CL, Dye AE, Díaz-Muñoz SL. 2023. Open questions in the social lives of viruses. Journal of Evolutionary Biology 36:1551-1567.
- Brunialti M, Höfler T, Nascimento M, Trimpert J. 2023. Suicidal Phenotype of Proofreading-Deficient Herpes Simplex Virus 1 Polymerase Mutants. J Virol doi:10.1128/jvi.01359-22:e0135922.
- Xing N, Höfler T, Hearn CJ, Nascimento M, Camps Paradell G, McMahon DP, Kunec D, Osterrieder N, Cheng HH, Trimpert J. 2022. Fast forwarding evolution—Accelerated adaptation in a proofreading deficient hypermutator herpesvirus. Virus Evolution doi:10.1093/ve/veac099.
- Nouailles G, Wyler E, Pennitz P, Postmus D, Vladimirova D, Kazmierski J, Pott F, Dietert K, Muelleder M, Farztdinov V, Obermayer B, Wienhold S-M, Andreotti S, Hoefler T, Sawitzki B, Drosten C, Sander LE, Suttorp N, Ralser M, Beule D, Gruber AD, Goffinet C, Landthaler M, Trimpert J, Witzenrath M. 2021. Temporal omics analysis in Syrian hamsters unravel cellular effector responses to moderate COVID-19. Nature Communications 12:4869.
- Trimpert J, Dietert K, Firsching TC, Ebert N, Thi Nhu Thao T, Vladimirova D, Kaufer S, Labroussaa F, Abdelgawad A, Conradie A, Höfler T, Adler JM, Bertzbach LD, Jores J, Gruber AD, Thiel V, Osterrieder N, Kunec D. 2021. Development of safe and highly protective live-attenuated SARS-CoV-2 vaccine candidates by genome recoding. Cell Rep 36:109493.
- Pennetzdorfer N, Höfler T, Wölflingseder M, Tutz S, Schild S, Reidl J. 2021. σ(E) controlled regulation of porin OmpU in Vibrio cholerae. Mol Microbiol 115:1244-1261.
- Lembke M, Höfler T, Walter A-N, Tutz S, Fengler V, Schild S, Reidl J. 2020. Host stimuli and operator binding sites controlling protein interactions between virulence master regulator ToxR and ToxS in Vibrio cholerae. Molecular Microbiology 114:262-278.
- Bischof K, Schiffer D, Trunk S, Höfler T, Hopfer A, Rechberger G, Koraimann G. 2020. Regulation of R1 Plasmid Transfer by H-NS, ArcA, TraJ, and DNA Sequence Elements. Frontiers in Microbiology Volume 11 - 2020.